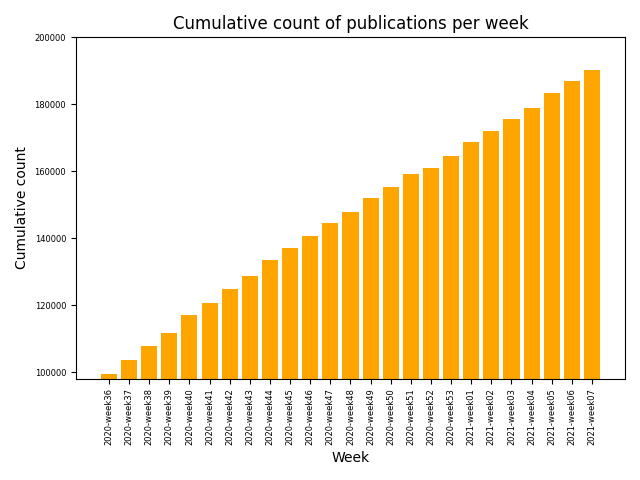
At present, SARS-CoV-2 and COVID-19 related literature is being published at an exponential rate, and the resulting flood of information is becoming effectively unmanageable by humans. Given the efforts of scientific community on identifying and structuring the relevant literature, increasing supply of COVID-19 related literature databases can be observed. Managing and extending these document collections – also called as corpora – in an automated fashion forms the foundation for further text mining applications.
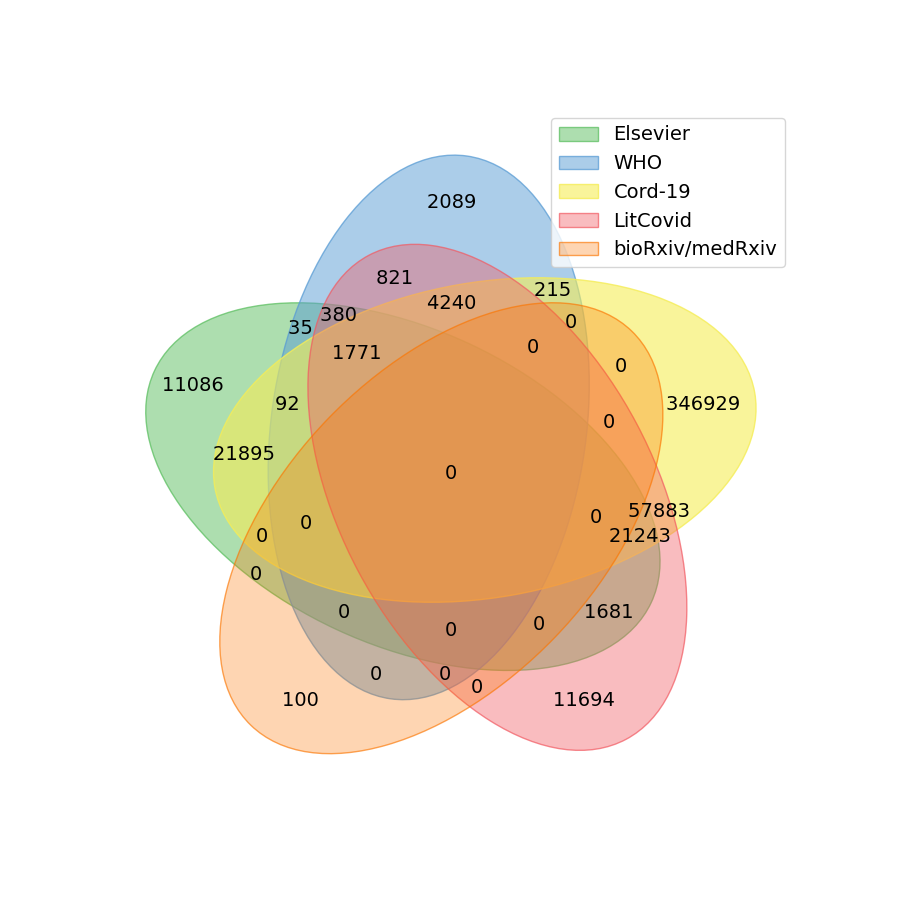
Currently, several organizations are collecting and regularly updating COVID-19 related corpora. These often include the textual document with metadata (such as publication dates, author list, PubMed IDs, DOIs). Some of the freely available corpora are listed below:
- LitCovid, provided by the National Institute of Health (NIH), USA., classifies manually the documents in classes such as general information, mechanisms, diagnosis, transmission.
- WHO COVID-19 corpus also categorizes the relevant literature.
- COVID-19 Open Research Dataset (CORD-19), created by the Allen Institute For AI et al., provides scientific literature for text miners and data scientists.
- COVID-19 related medRxiv/bioRxiv preprints gives scientists a platform to publish research without before peer-review
- Elsevier’s Coronavirus Information Center includes biological and medical research on the SARS-CoV-2 and COVID-19
Moreover, full-text is available for the Elsevier corpus, the medRxiv and bioRxiv preprints, as well as the CORD-19 corpus, which we integrate in our literature mining platform SCAIView. Other noteworthy resources include the Dimensions literature database and the Cochrane special collections.
Based on the available abstracts provided in the merged databases, unsupervised topic modelling approaches such as Latent Dirichlet Allocation (LDA) provide insights into different subjects covered by the corpora. Additionally, by using topic tags provided in, for instance, the LitCovid corpus, it is also possible to train a document classifier that can be used to categorize new COVID-19 related literature. In the near future, the aim is to continuously integrate further resources and more full-text, thereby validating and extending the existing text mining approaches.